
Parts
We are nominating our part collection for the Best Part Collection Award.
We are nominating our BBa_252HSV3T part for the Best New Composite Part Award.
Overview
The purpose of this parts collection is to enable modular engineering of Acinetobacter baylyi ADP1 for applications in environmental DNA (eDNA) sensing and general chromosomal modification. By exploiting ADP1's natural competence2 and homologous recombination machinery, these parts allow for the creation of customizable biosensors and their integration into ADP1's chromosome. These sensors function to detect specific DNA sequences from environmental samples.
Our team focused on designing components to support a recombination-based detection system to target eDNA from harmful algal blooms (HABs), specifically sequences associated with microcystin production. However, the modular design allows the target sequence to be easily substituted, enabling adaptation to other genetic targets or environmental biomarkers. This parts collection can be used by others to make their own eDNA sensors or engineer the ADP1 genome.
Circuit Design
Briefly, the biosensor circuit employs two sets homology arms to facilitate targeted DNA exchange between the ADP1 genome and incoming environmental sequences (see Figure 1).
- 1. Outer homology arms (integration set): Homologous to sequences that flank the ACIAD2049 locus in ADP1, which was previously characterized as a non-essential genomic site by the Austin-UTexas 2022 team. This enables stable chromosomal integration of the detection circuit.
- 2. Inner homology arms (target recognition set): Designed to recognize a highly conserved region within the mcy gene cluster of M. aeruginosa. These flanks enable recombination when the ADP1 chassis encounters matching mcy-derived eDNA.
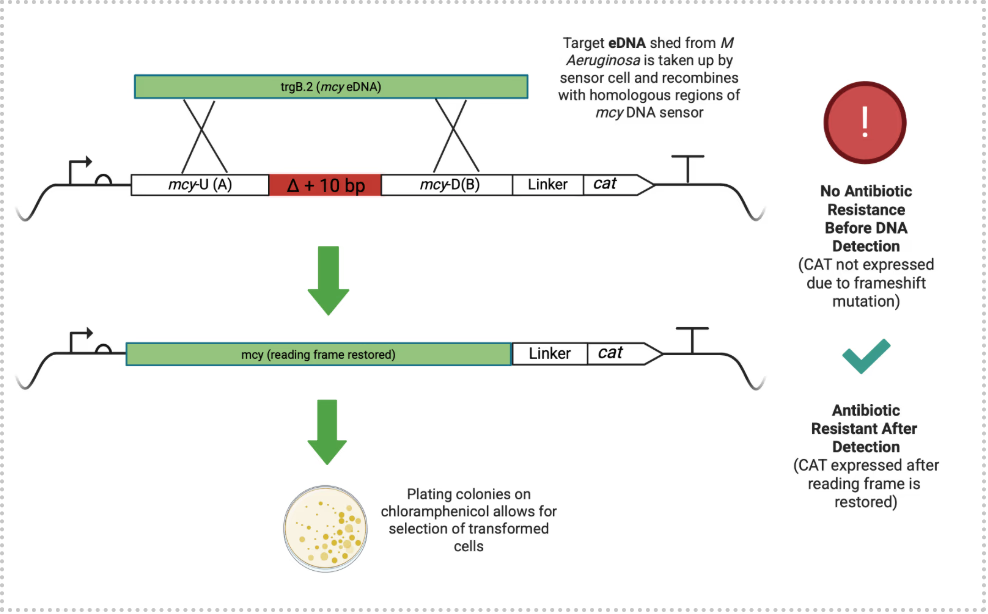
Figure 1. Mechanism of Recombination-Based Sensing. The chromosomally integrated sensor contains a frameshifted chloramphenicol resistance (CAMr) gene. Upon encountering homologous mcy eDNA, a recombination event restores the correct reading frame, enabling cell growth on media containing chloramphenicol (CAM).
Part Categories
The collection includes several broad categories of parts which collectively serve as a modular system of basic and composite parts for incorporating an mcy eDNA sensor in ADP1:
- Homology flanks (6 parts) for targeted ADP1 chromosomal integration and recombination-based sensing, including the ACIAD2049 upstream/downstream flanks, ACIAD0611 upstream/downstream flanks, and mcyB target sequence upstream/downstream regions.
- Sensor/Reporter Components (2 parts) that will comprise sensor composite parts, including the CP25 constitutive promoter and chloramphenicol resistance gene with protein linker.
- Sensor Composite Parts (3 parts) comprised of more than one of the above sensor/reporter components: mcyB 3 in 1 sensor, mcyB sensor region with 10 base pair insertion, mcyB 3-in-1 target, mcyB Sensor with ACIAD2049 Homology Flanks from Acinetobacter baylyi ADP1-ISx.4
- Custom Backbone Composite Parts (4 parts) for facilitating Golden Gate Assembly (GGA) for modular part construction and ADP1 chromosomal insertion.
- Recombinase System (4 parts) for enhancing recombination efficiency of shorter homology flanks in ADP1. Includes parts from the Acinetobacter baumanii recombinase RecAB system.1
- Target DNAs (7 parts) for testing sensor and recombinase constructs.
Together, these elements form a standardized toolkit for designing new eDNA biosensors or integrating genetic constructs in ADP1. See parts sheet below breakdown of all parts in our team's parts collection.
Golden Gate Assembly Strategy
Basic parts are first cloned into plasmid backbones, followed by composite part assembly into complete sensor constructs (see Figure 1). This hierarchical and standardized format allows rapid reassembly of new sensor variants by swapping in alternate target or reporter sequences. All part-type overhangs follow the Bee Toolkit/Yeast Toolkit framework.3,6
- Each DNA sequence was PCR amplified, with the addition of BsaI restriction sites and part-type specific overhangs.
- PCR amplification also flanked the BsaI cut sites and overhangs with BsmBI sites, enabling ligation into plasmid backbones.
- Note that for some parts, such as the basic parts that comprise the ACIAD custom backbones, the sequences were ordered with the cut sites and part-type specific overhangs already included. Instead of BsaI, PaqCI restriction sites were present.
- Basic parts were removed from their respective plasmid backbone through GGA with BsaI digestion, leaving specific overhangs that guide the assembly order.
- Promoters, coding sequences, terminators, and homology flanks were combined into either integration or rescue cassettes (more on these categories below).
- Note that for some parts, such as the basic parts that comprise the ACIAD custom backbones, composite part assembly was driven by PaqCI digestion/ligation.
Phase 1 Basic Part Construction:
Phase 2 Composite Part Construction:
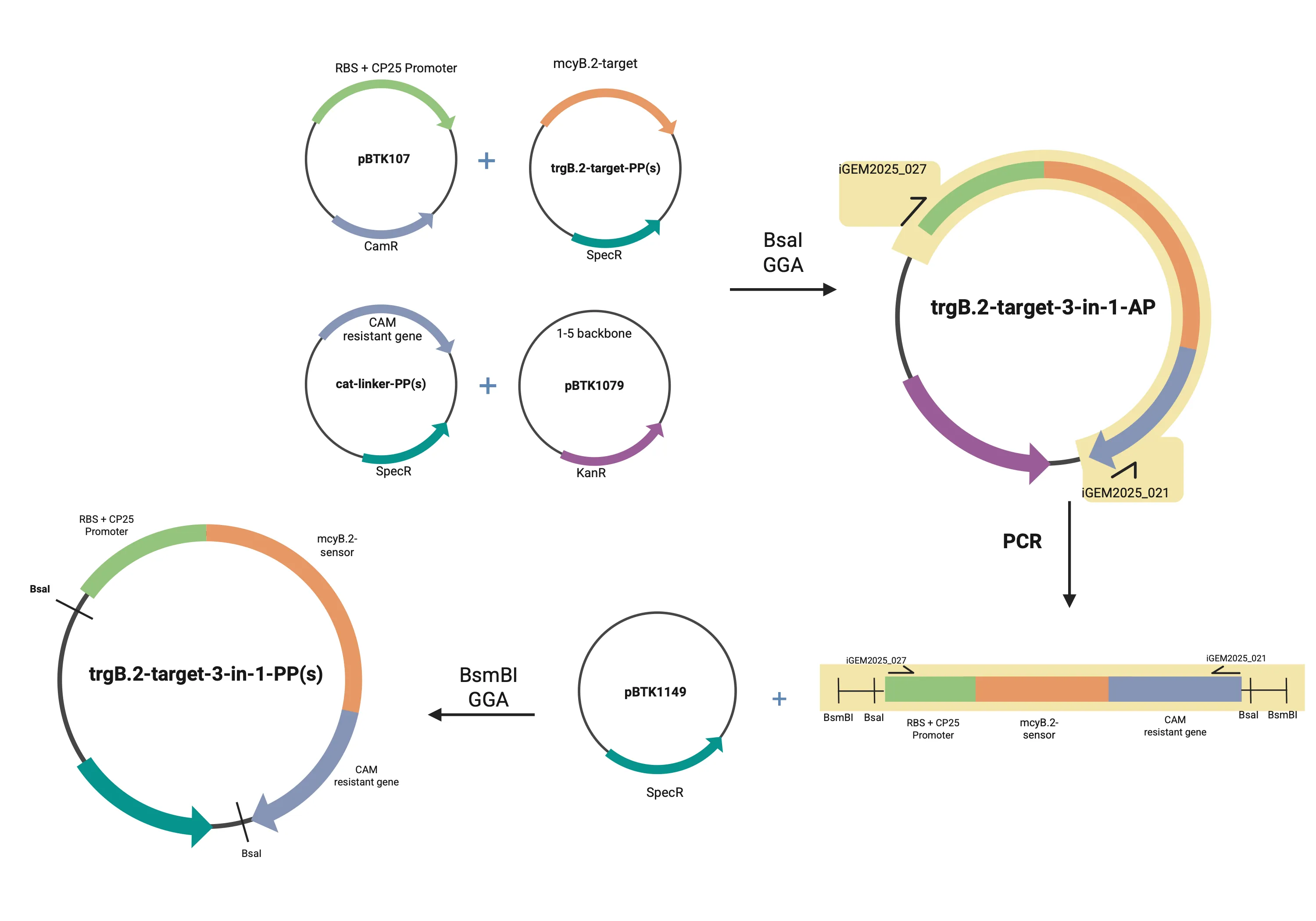
Figure 2. Shows process of creating the mcy eDNA sensor composite part. The top left shows part plasmids being created from the basic parts using BsmBI digestion-ligation in phase I. In phase II, through BsaI digestion-ligation, the basic parts were ligated in into a backbone to create an assembly plasmid.
ADP1 Genetic Engineering Strategy
The following engineering workflow was derived from the Austin-UTexas 2022 iGEM team.5
- ADP1 undergoes a transformation reaction with the composite part containing the tdk (azidothymidine susceptibility) and kan (kanamycin resistance) genes flanked by ACIAD2049 homology arms. After recombination, the ACIAD2049 gene in ADP1 should be replaced with the tdk/kan cassette.5
- Successful transformants will be selected for using kanamycin (kan) plates. In practice, this means cells successfully transformed with the cassette will demonstrate visible growth on LB agar-kan plates due to the resistance conferred by the kanR gene.
- ADP1 is transformed with a composite part flanked by ACIAD2049 homology arms to replace the tdk/kan cassette.
- Successful transformants will be selected for using LB agar-azidothymidine (AZT) plates. This is because the tdk/kan cassette makes ADP1 susceptible to AZT. Removing the cassette from the genome removes this susceptibility brought on by the tdk gene.
Incorporation of tdk/kan cassette:
Rescue and Insertion:
Computational Design
Various software was used to identify conserved target regions were across multiple genomes and verify that unwanted transcriptional or translational artifacts were minimized. For more details on computational validation, see the link to the mcyB sensor region with 10 base pair insertion: BBa_25T8GWZR.
Prize Nominations and Narratives
Our team nominates our parts collection for the best parts collection special prize. The CyanoSense parts collection establishes a modular framework that enables Acinetobacter baylyi ADP1 to be rapidly engineered for diverse applications, from detecting environmental toxins to performing targeted genomic edits. By integrating a recombination-based sensing mechanism with standardized Golden Gate backbones containing homology flanks and regulatory elements, the system streamlines circuit assembly and broadens ADP1's utility as a chassis for recombination-driven biosensing and genome engineering.
Furthermore, our team nominates our mcyB Sensor with ACIAD2049 Homology Flanks from Acinetobacter baylyi ADP1-ISx4 for the best composite part special prize. This composite part offers a modular platform for DNA-based sensing. Because ADP1 is naturally competent, this chassis enables the detection of environmental DNA sequences via transformation without external reagents or cell-free systems. By modifying the target sequence within the frameshifted region, teams could rapidly adapt the sensor to detect a wide variety of genes, allowing for use in a variety of potential sensing contexts, from the environment to healthcare. The use of chromosomal integration, rather than plasmid-based expression, improves biosafety and long-term stability, which are key considerations for environmental deployment.
Submitted Basic and Composite Parts
References
- Tucker AT, Nowicki EM, Boll JM, Knauf GA, Burdis NC, Trent MS, Davies BW. Defining gene-phenotype relationships in Acinetobacter baumannii through one-step chromosomal gene inactivation. mBio. 2014 Aug 5;5(4):e01313-14. doi:10.1128/mBio.01313-14. PMID:25096877. PubMed
- Santala S, Santala V. Acinetobacter baylyi ADP1-naturally competent for synthetic biology. Essays Biochem. 2021 Jul 26;65(2):309-318. doi:10.1042/EBC20200136. PMID:33769448. Science+3ACS Publications+3PMC+3
- Leonard SP, Perutka J, Powell JE, Geng P, Richhart DD, Byrom M, Kar S, Davies BW, Ellington AD, Moran NA, Barrick JE. Genetic Engineering of Bee Gut Microbiome Bacteria with a Toolkit for Modular Assembly of Broad-Host-Range Plasmids. ACS Synth Biol. 2018 May 18;7(5):1279-1290. doi:10.1021/acssynbio.7b00399. PMID:29608282. ACS Publications+2PMC+2
- Chuong J, Brown KW, Gifford I, Mishler DM, Barrick JE. Engineered Acinetobacter baylyi ADP1-ISx Cells Are Sensitive DNA Biosensors for Antibiotic Resistance Genes and a Fungal Pathogen of Bats. ACS Synth Biol. 2025 Jul 18;14(7):2488-2493. doi:10.1021/acssynbio.5c00360. PMID:40579381. PubMed
- 2022 UT Austin iGEM Team. Part Collection. In: 2022 UT Austin iGEM Wiki. Austin, TX: The University of Texas at Austin; 2022. Available from: https://2022.igem.wiki/austin-utexas/parts
- Lee ME, DeLoache W, Cervantes B, Dueber JE. A Highly Characterized Yeast Toolkit for Modular, Multipart Assembly. ACS Synth Biol. 2015 Sep 18;4(9):975-986. doi:10.1021/sb500366v. PMID:25871405.
