
Contribution
Introduction
Throughout the course of this project, our team was eager to contribute to the development of the vast field of synthetic biology and future iGEM teams. Our contribution focuses on expanding the toolbox for engineering Acinetobacter baylyi (ADP1), a powerful but still underutilized synthetic biology chassis. We created new plasmid backbones, standardized assembly parts, and modular biosensor designs that will enable future iGEM teams to edit A. baylyi ADP1 more efficiently and to adapt it for diverse applications. Our biological parts are well-characterized and documented on our team's Parts Page and the iGEM Parts Registry.
Backbone and Plasmid Design for ADP1 Genome Engineering
Our team's major contribution to iGEM and to future teams lies in the generation of novel parts for advancing work with ADP1. Building on the work of the 2022 UT Austin iGEM team, who demonstrated targeted integration into ADP1 using homology flanks,1 we expanded on the functionality of their modular system.
- We created paired upstream (5') and downstream (3') homology flank libraries for two well-characterized loci (ACIAD0611 and ACIAD2049). These flanks contain BsmBI or PaqCI recognition sites and unique overhangs, enabling seamless assembly of functional plasmids with promoters, coding sequences, and terminators.2 See our Parts Page for important DNA sequences and Experiments Page for Acinetobacter baylyi genetic engineering protocols.
- We engineered a ColE1/AmpR backbone with built-in ADP1 homology flanks and a GFP dropout cassette, enabling Golden Gate-based insertion of custom genetic modules, such as promoters, enzymes, and reporter genes, into the ADP1 chromosome with fewer cloning steps.
This modular system reduces the time and complexity required for ADP1 integration, providing future teams with a ready-to-use backbone for targeted additions, deletions, and edits without needing to re-develop basic tools.
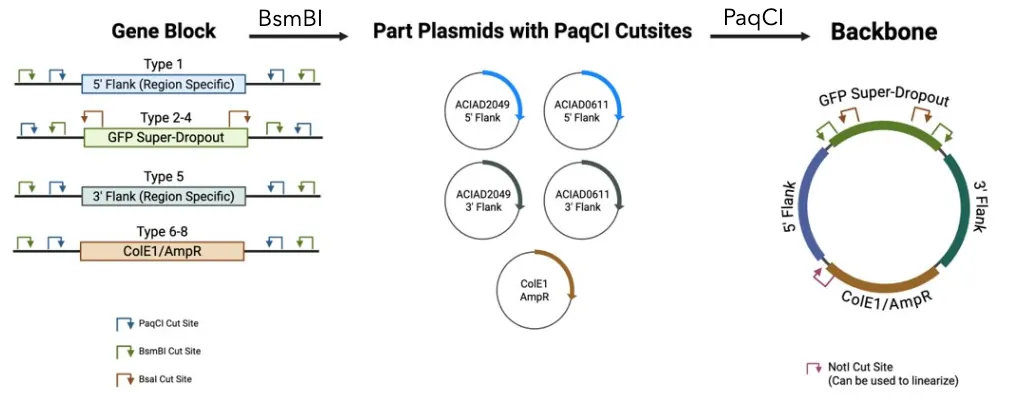
Figure 1. Process for engineering a modular novel Acinetobacter baylyi ADP1 backbone using PaqCI Sites and Homology Flanks. Four gBlocks (gene blocks) from IDT were ordered. Using BsmBI on the 5' Flank gBlock, 3' Flank gBlock, and the ColE1 and AmpR gBlock, a part plasmid was assembled with a pBTK1001 backbone. Using these part plasmids and the GFP Super-Dropout gBlock, a backbone plasmid will be assembled.
Expanding ADP1 as a Synthetic Biology Chassis
While the previous 2022 iGEM team focused on small edits, we aimed to test Acinetobacter baylyi (ADP1's) capacity for functionally diverse genetic integrations through a modular backbone construct and integrating methods of increasing its recombination capabilities.
With these goals in mind, we created a series of designs to improve the functionality of ADP1 for use by future teams:
- Designed a modular integration framework with standardized flanks, promoters, and terminators.
- Designed and tested a system using the well-characterized recombinase enzyme to increase the bacterium's recombination frequency
See our Parts Page, Results Page, and Engineering Success Page for more details.
Even where integrations were incomplete, we provide tested designs, protocols, and lessons learned. By charting both our successes and limitations, we provide a roadmap for teams interested in pushing ADP1 towards more complex biological engineering.
Engineering a DNA Biosensor for M. aeruginosa eDNA Detection
In our project, we developed a DNA biosensor and integrated it into the ADP1 chromosome, designing a system that can be adapted to detect a variety of eDNA targets.3,4 To support future projects, our team created several plasmid parts, including ADP1 homology flanks, promoters, and terminator sequences, which can be reused for the development of new eDNA sensors. Our modular design enables straightforward incorporation of new Type 3 parts corresponding to conserved DNA sequences for specific targeting. See our development process in Inserting the DNA Sensor into ADP1 Chromosome on the Engineering Success Page and detailed outcomes on our Results Page. This modular approach will help facilitate a generation of customized DNA biosensors for a wide range of applications.
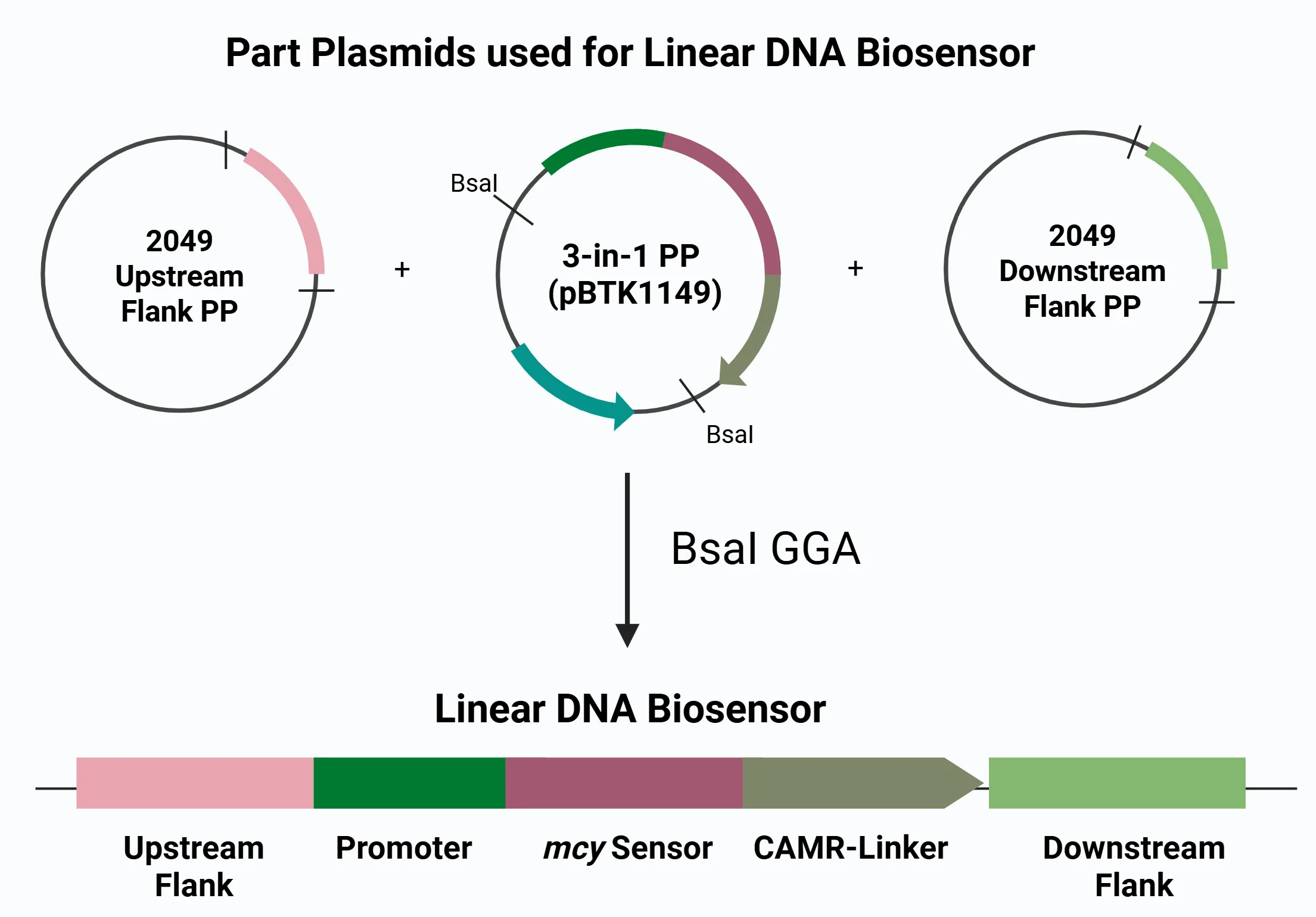
Figure 2. Modular microcystin biosensor construct. A 5' ACIAD2049 upstream flank, a 3' ACIAD2049 downstream flank, and a 3-in-1 part plasmid (pBTK1149 backbone) are joined through Golden Gate Assembly. The result is a linear product containing a promoter-driven microcystin sensor flanked by homology regions and a CAMR-linker for selection. This strategy enables precise and efficient integration of biosensor modules into Acinetobacter baylyi (ADP1's) genome for the detection of microcystin DNA.
Additionally, we explored a novel approach to use a recombinase plasmid to increase our biosensor's recombination frequency with shorter homology flanks. This applies our sensor to more realistic environmental conditions, as the target eDNA can degrade over time and be harder to detect. Together, these contributions not only expand the toolbox for DNA sensing but also open up new possibilities for designing flexible and efficient biosensors in synthetic biology.
Proof of Concept Tests for DNA Biosensor
We successfully demonstrated that ADP1 can integrate a DNA biosensor into its chromosome and respond to environmental DNA (eDNA) with measurable outputs (Engineering Success). This provides the foundation for developing modular biosensors that detect harmful genes in real-world contexts.
Key contributions include:
- Chromosomal Integration: We confirmed that our biosensor constructs were stably integrated into the Acinetobacter baylyi (ADP1) genome using the tdk/kan cassette's selective properties, validating ADP1 as a reliable host for synthetic sensors. This process also highlights the usage of selective markers to qualitatively confirm results.
- Detection Thresholds: We tested sensor performance across different concentrations of target part plasmid DNA, generating a baseline understanding of detection limits for ADP1. This provides reference for future teams if they test ADP1 sensor detection limits with different DNA concentrations.
- Specificity Testing: We evaluated sensor response to genomic DNA (gDNA) versus non-target DNA, showing that the biosensor can distinguish between specific sequences. We also began initial tests of our sensor in mixed media to apply a more realistic condition for the DNA biosensor. These tests can be easily replicated from our documented protocols (Experiments Page) for future eDNA sensors in ADP1.
Together, these results represent an early but important step toward building field-deployable biosensors. While our focus was on microcystin-producing genes, the modular design means the same approach could be extended to other environmental DNA targets for design, integration, and testing.
References
- UTiGEM22 wiki. (n.d.). https://2022.igem.wiki/austin-utexas/index
- Engler, C., Kandzia, R., & Marillonnet, S. (2008). A One Pot, One Step, Precision Cloning Method with High Throughput Capability. PLoS ONE, 3(11), e3647. https://doi.org/10.1371/journal.pone.0003647
- Chuong, J., Brown, K. W., Gifford, I., Mishler, D. M., & Barrick, J. E. (2025). Engineered Acinetobacter baylyi ADP1-ISx Cells Are Sensitive DNA Biosensors for Antibiotic Resistance Genes and a Fungal Pathogen of Bats. ACS Synthetic Biology. https://doi.org/10.1021/acssynbio.5c00360
- Rizzi, A., Pontiroli, A., Brusetti, L., Borin, S., Sorlini, C., Abruzzese, A., Sacchi, G. A., Vogel, T. M., Simonet, P., Bazzicalupo, M., Nielsen, K. M., Monier, J., & Daffonchio, D. (2007). Strategy for in situ detection of Natural Transformation-Based Horizontal Gene Transfer Events. Applied and Environmental Microbiology, 74(4), 1250-1254. https://doi.org/10.1128/aem.02185-07
